MIT researchers use AI to find drugs that could be repurposed for COVID-19
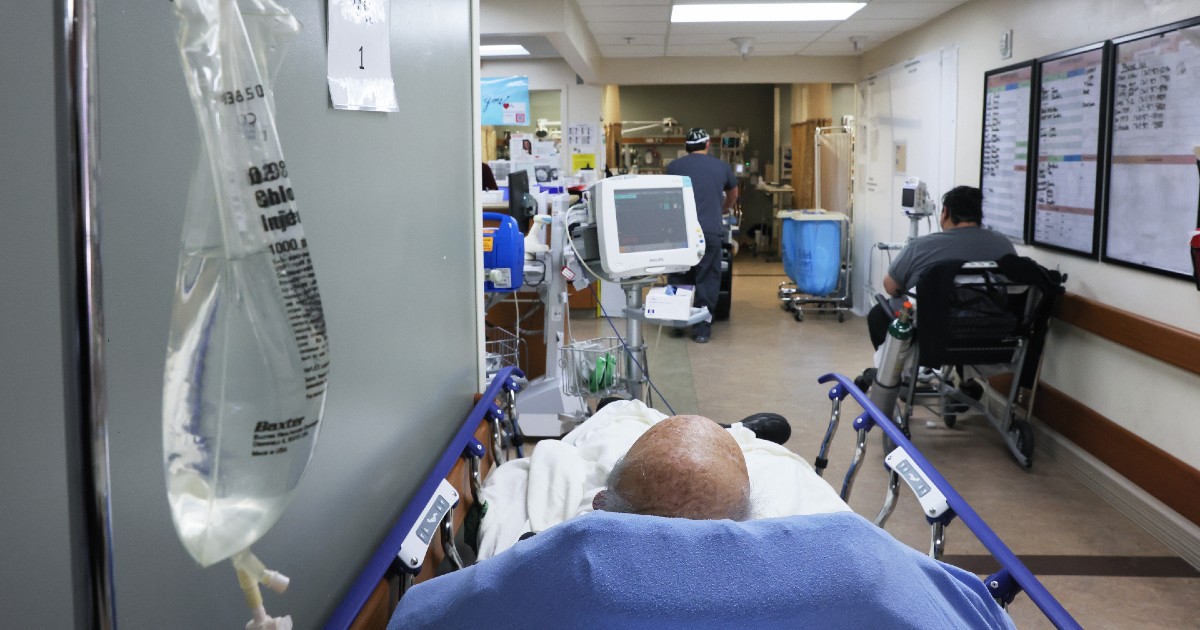
(Photo by Mario Tama/Getty Images)
The Massachusetts Institute of Technology announced this week that researchers had used machine learning to identify medications that may be repurposed to fight COVID-19.
"Making new drugs takes forever," Caroline Uhler, a computational biologist in MIT's Department of Electrical Engineering and Computer Science and the Institute for Data, Systems and Society, said in a press statement. "Really, the only expedient option is to repurpose existing drugs."
The research from Uhler's team, which appears in the journal Nature Communications, notes that the novel coronavirus tends to have much more severe effects in older patients.
"Since the mechanical properties of the lung tissue change with aging, this led us to hypothesize an interplay between viral infection/replication and tissue aging," wrote the researchers.
WHY IT MATTERS
The researchers pointed out that lung tissue becomes stiffer as a person gets older, and it shows different patterns of gene expression than in younger people in response to the same signal.
"We need to look at aging together with SARS-CoV-2 – what are the genes at the intersection of these two pathways?" said Uhler.
As the study explains, the team generated a list of possible drugs using an autoencoder before mapping the network of genes and proteins involved in aging and novel coronavirus infection. They then pinpointed genes causing cascading effects throughout the network using statistical algorithms.
"Among the various protein kinases ... identified by our drug repurposing pipeline, RIPK1 was singled out by our causal analysis as being upstream of the largest number of genes that were differentially expressed by SARS-CoV-2 infection and aging," wrote the researchers in the study. In other words, drugs that act on RIPK1 may have the potential to treat COVID-19.
"Given the distinct pathways elicited by RIPK1, there is a need to develop appropriate cell culture models that can differentiate between young and aging tissues to validate our findings experimentally and allow for highly specific and targeted drug discovery programs," read the study.
THE LARGER TREND
Machine learning and artificial intelligence have been instrumental for many facets of COVID-19 research, with scientists using them to predict the length of hospitalization and probable outcomes among patients, as well as to detect the disease in lung scans and improve treatment options.
Cris Ross, CIO at the Mayo Clinic, said in December that AI has been key to understanding COVID-19.
Around the world, Ross said, algorithms are being used to "find powerful things that help us diagnose, manage and treat this disease, to watch its spread, to understand where it's coming next, to understand the characteristics around the disease and to develop new therapies."
ON THE RECORD
"While our work identified particular drugs and drug targets in the context of COVID-19, our computational platform is applicable well beyond SARS-CoV-2, and we believe that the integration of transcriptional, proteomic, and structural data with network models into a causal framework is an important addition to current drug discovery pipelines," wrote the MIT research team.
The Diverse Roles of AI & ML
Machine learning applications are proliferating and improving, and this month we'll explore the many ways they're being put to work.
Kat Jercich is senior editor of Healthcare IT News.
Twitter: @kjercich
Email: kjercich@himss.org
Healthcare IT News is a HIMSS Media publication.